
Computational pathology uses the power of deep machine learning, image analytics and big data integration to enhance diagnostic precision and transform the next generation of pathologists.
Philips is leading the way in computational pathology research, delivering novel technologies for image analytics, which are fully embedded within their digital pathology scanning, storage and display solution.
Computational Pathology Empowering Pathologists to Support Improved Patient Care
We're in a really exciting time in pathology
Need for computational pathology 1
We are moving rapidly into an era where next-generation pathology is becoming a reality with the advent of digital pathology. Paradigm shifts are being witnessed in cancer care with precision medicine and personalized treatments increasing by the day.
Pathologists are absolutely central to this dream of personalized medicine. It is at the desk of the pathologist that first clinical decisions regarding the patient are made and that will continue to be the case.
Increasing workload 2
70%

projected increase in new cancer cases within the next two decades
20%

of pathologists work overtime weekly or have to outsource services
Shortage of pathologists 3
60%

of active physicians in pathology are age 55 or older
10%

decrease in active physicians in pathology in 2008 - 2013
Expert opinions
A transformation in computational pathology 4
The evolution of deep learning and the improvement in accuracy for image pattern recognition has been staggering in the last few years. Everything from bio-metric and security, voice recognition and intelligent advertising, to driver-less cars is powered by deep learning technologies.
Next up is pathology. We believe that digital pathology and machine learning combined, could empower pathologists with new tools, and help drive improvements in workflow and diagnostic precision in both discovery and diagnosis.
Recent convergence of technologies, together with Philips innovation in computational pathology will help us to transform pathology together.
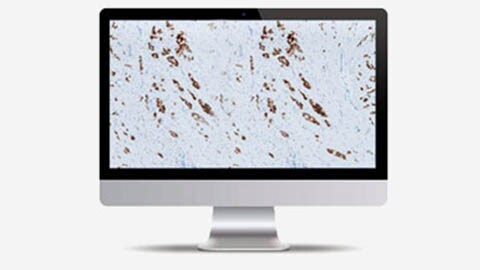
Clinical algorithms5
We have an extensive roadmap for the delivery of CE-IVD approved algorithms to support clinical decision making in multiple tissues. Initially we are providing digital image analysis applications in support of the pathologist for ER, PR, HER2 and Ki-67 clinical assessments.
Resources
Related articles



